… at least from my perspective. I’ll now stop ranting about the hype and media coverage of ENOCDE, and extend my compliments to the consortium for an amazingly well-coordinated effort to achieve an impressive level of consistency and quality for such a large consortium. Whatever else you might want to say about the idea of ENCODE, you cannot say that ENCODE was poorly executed.
It’s time to get into the interesting stuff – what’s actually in the papers. Among the results I’ve been most eagerly awaiting to see in print are the DNase hypersensitivity results now published in Thurman et al. (Nature 489, 75–82 (06 September 2012) doi:10.1038/nature11232)
Why is this interesting? Because it raises provocative and possibly disturbing questions regarding how transcription factors navigate and read out information from the genome.
Most interesting to me are the results you can see in Figure 2A, which I’ve cropped and pasted below:
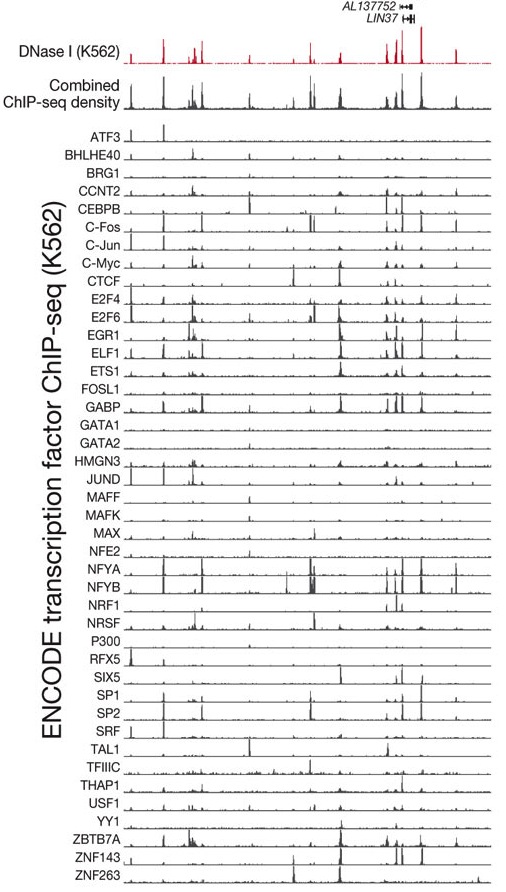
In the top trace of the figure you see sites of accessible chromatin as indicated by DNase hypersensitivity. In the row below it, you see where transcription factors (TFs) bind, in aggregate. Clearly, TFs bind to regions of accessible chromatin – no surprise there.
But now look at the binding traces for individual TFs: the shocker is that binding at accessible DNA doesn’t seem to be very specific. In any open region, you find lots of binding by many different TFs. This suggests either that regulation is highly complex, requiring dozens of TFs at each open site, or that TF binding is extremely promiscuous and messy.
I’m not a believer in complexity. I believe in robust simplicity embedded in messiness. The nucleus is stuffed with biochemically active players, and interactions are inevitably promiscuous, creating a messy interaction network. There’s no way to prevent the messiness, because negative selection against it isn’t strong and genetic drift is real. The genome is more like a jungle than a finely tuned watch mechanism. Transcriptional regulation works in spite of the mess, not because of complexity.
And so the questions we really need to address are: What is the logic that underlies gene regulation? And how does this regulatory logic remain robust within its messy context? My position is that we really don’t know how the signal gets sorted from the noise in the genome, and that our traditional models of how transcription factors find their targets and regulate transcription have serious problems. And right now I’m in possession of some new, unpublished data that really makes this problem disturbing. Stay tuned…

Thanks Mike, that was pretty interesting, and I was able to follow your analysis once I suspended my ignorance of the basics. But I still would love if you’d explain some basics: what do TFs do in the world as I experience it? And why does the messiness matter in that world?
Great question – I think we need a post that explains what TFs do, what exactly the ENCODE scientists measured, and why we expect these measurements to tell us something important.
The quick answer is that everything in life (almost 😉 ) depends on having the right genes switched on or off at the right time and place. TFs are proteins that make that happen by binding to specific places in the genome and recruiting the cell’s gene expression machinery to genes.
But to do their job, TFs have to find the correct spot to bind in a very large genome. TFs aren’t very specific – they recognize very short DNA sequences (6-10 nucleotides generally, in a 3 billion nucleotide genome). This is one source of messiness that the cell has to deal with.
So a key question for us is, how do we identify those important (i.e. functional) binding sites in this large genome? Various ENCODE experiments address that question – by measuring directly where TFs bind, and by measuring where the genome is essentially open and accessible to TFs in search of their binding sites. Accessible DNA is what the DNase hypersensitivity experiments identify.
Clarification: the relationship between accessible DNA and TF binding is one of the questions here – is accessibility necessary in order for TFs to bind, or do TFs induce accessibility? Some of both clearly happens.